Arabidopsis Research Roundup: December 18th
This festive Arabidopsis Research Roundup begins with a commentary article from a global consortium of plant scientists who propose a framework of future training for researchers who will take advantage of the experimental tools available in Arabidopsis. Secondly is study from Caroline Dean (JIC) that defines the role of the LHP1 protein in epigenetic control of gene expression. Thirdly John Doonan (Aberystwyth) is a co-author of work that defines an important component of mitotic spindle formation. Next is a study led by Zinnia Gonzalez-Carranza in Nottingham that offers further insights into the function of the HWS gene. The fifth study comes from the lab of Alexander Ruban (QMUL), further investigating the importance of NPQ in photosynthetic control. The sixth paper from the Van Ooijen lab (Edinburgh) characterises the role of sumoylation in the control of CCA1 activity. The penultimate paper from the Harberd lab in Oxford defines the importance of DNA mismatch repair on genome sequence integrity whilst the final paper characterises the next phase in the long story of Arabidopsis ALF4 function and includes Charles Melynk (SLCU) as a co-author.
Friesner J et al (2017) The Next Generation of Training for Arabidopsis Researchers: Bioinformatics and Quantitative Biology. Plant Physiol. doi: 10.1104/pp.17.01490. Open Access

The current GARNet PI Jim Murray and past GARNet coordinator Ruth Bastow are authors in this international consortium that suggests future directions for the global Arabidopsis community. This consortium is led by Joanna Friesner and concludes that it is critical that the next generation of plant scientists receive appropriate training in bioinformatics and quantitative biology so as to take advantage of the remarkable array of datasets that are now available to Arabidopsis researchers.
Berry S, Rosa S, Howard M, Bühler M, Dean C (2017) Disruption of an RNA-binding hinge region abolishes LHP1-mediated epigenetic repression Genes Dev. doi: 10.1101/gad.305227.117 Open Access
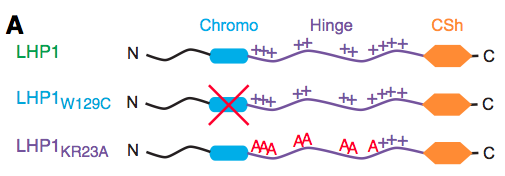
Caroline Dean (John Innes Centre) leads this study that investigates the role of the polycomb associated protein LIKE HETEROCHROMATIN PROTEIN 1 (LHP1) in the regulation of the repressive histone mark H3K27me3. They demonstrate that the intrinsically disordered hinge region of LHP1 is responsible for RNA-binding and that disruption of this region prevents the formation of sub-nuclei foci, provides a potential link to wider epigenetic regulation.
Lee YJ, Hiwatashi Y, Hotta T, Xie T, Doonan JH, Liu B (2017) The Mitotic Function of Augmin Is Dependent on Its Microtubule-Associated Protein Subunit EDE1 in Arabidopsis thaliana. Current Biol. doi: 10.1016/j.cub.2017.11.030
Open Access

John Doonan and colleagues at Aberystwyth University are co-authors on this study regarding the role of the Microtubule-Associated Protein Subunit EDE1, which is a member of the Augmin complex, during mitosis. EDE1 specifically localised with the augmin complex during spindle formation, a role that cannot be replaced by the homologous protein AUG8. This work reveals that specificity of the augmin complex can be determined by interaction with subunits that only contribute to complex function during particular phases of the cell cycle.
Zhang X, Jayaweera D, Peters JL, Szecsi J, Bendahmane M, Roberts JA, González-Carranza ZH (2017) The Arabidopsis thaliana F-box gene HAWAIIAN SKIRT is a new player in the microRNA pathway. PLoS One. doi: 10.1371/journal.pone.0189788 Open Access
Zinnia Gonzalez-Carranza (Nottingham) is the corresponding author on this study that follows on from work published earlier in 2017 regarding the role of the HAWAIIAN SKIRT gene is plant development. In this latest work they identify mutations in the previously characterized Exportin-5 HASTY gene as suppressors of the hws mutant phenotype. Further investigation shows that HWS genetically interacts with other genes involved in miRNA pathway indicates that HWS somehow interacts with biogenesis, accumulation or function of these small RNAs.
Townsend AJ1, Ware MA1, Ruban AV (2017) Dynamic interplay between photodamage and photoprotection in photosystem II. Plant Cell Environ doi: 10.1111/pce.13107
In this paper Alexander Ruban (QMUL) is the corresponding author on work that expands his groups contribution to the understanding of the role non-photochemical quenching (NPQ) plays during photoinhibition. In this work they compare the activity of NPQ versus endogenous photosystemI repair mechanisms in the maintenance of photosynthetic activity during photoinhibitory conditions. Overall they conclude that NPQ is a more important mechanism for photoprotection under short periods of illumination.
Hansen LL, Imrie L, Le Bihan T, van den Burg HA, van Ooijen G (2017) Sumoylation of the Plant Clock Transcription Factor CCA1 Suppresses DNA Binding. J Biol Rhythms doi: 10.1177/0748730417737695 Open Access
This paper from the Van Ooijen lab accompanies one that was featured in last weeks ARR and extends their finding that sumoylation plays an important role in control of the circadian clock. In this paper they show that the CCA1 clock protein is sumoylated and that perturbing this modification alters the binding of CCA1 to a target promotor, even though it’s localization or stability were unaffected. Using an in vitro system they show that sumoylation is a direct determinant of CCA1 binding to its target promotor suggesting that this PTM fine tunes the activity of this key circadian control element.
Belfield EJ, Ding ZJ, Jamieson FJC, Visscher AM, Zheng SJ, Mithani A, Harberd NP (2017) DNA mismatch repair preferentially protects genes from mutation. Genome Res. doi: 10.1101/gr.219303.116
Past GARNet Advisory board member Nick Harberd (Oxford) leads this multi-generational study on the effect of DNA mismatch repair (MMR) on maintenance of an entire genome. They perform whole genome sequencing across five generations of Arabidopsis plants with a mutation in the MMR pathway and show that particular types of nucleotide error are more prevelant amongst the total 9000 mutations that accumulate. Interestingly they show that single nucleotide variants are more likely to accumulate in genic regions, indicating that protein coding areas of the genome are preferentially protected from damage.
Bagchi R, Melnyk CW, Christ G, Winkler M,, Kirchsteiner K, Salehin M, Mergner J, Niemeyer M, Schwechheimer C, Calderón Villalobos LIA, Estelle M (2017) The Arabidopsis ALF4 protein is a regulator of SCF E3 ligases. EMBO J. doi: 10.15252/embj.201797159
During his time as a research fellow at the Sainsbury lab in Cambridge. Charles Melynk contributed to this research that is a throwback to the early day of Arabidopsis mutant analysis. The alf4 was first described as a possible auxin mutant in 1995 and this work brings this study full circle by characterising the ALF4 protein as a novel regulator of SCF complexes, which are known to be involved in auxin and GA signaling. ALF4 specifically functions by interacting with the SCF-core component RBX1. Future work will determine whether this effect is specific to SCFs involved in hormone signaling or whether it is a more general effect.

Leave a Reply