This weeks Arabidopsis Research Roundup begins with two papers that look at endogenous and exogenous causes of cell proliferation. Firstly Mike Bevan (JIC) leads a team that looks into the role of controlled protein degradation in this process whilst secondly, Peter Etchells from Durham is a co-author on a study that investigates how nematode pathogens stimulate cell proliferation at the site of infection.
Thirdly is work featuring Cyril Zipfel and colleagues from TSL that looks at how autophosphorylation controls the activity of calcium dependent protein kinases. Fourthly is a broad collaboration led by Richard Mott (UCL) that uses genomic structural variation to identify novel loci. Next Simon Turner from the University of Manchester phylogenetically defines the RALK peptide lineages across plant species. Finally researchers at the University of York conduct a structural analysis of the Arabidopsis AtGSTF2 glutathione transferase.
Dong H, Dumenil J, Lu FH, Na L, Vanhaeren H, Naumann C, Klecker M, Prior R, Smith C, McKenzie N, Saalbach G, Chen L, Xia T, Gonzalez N, Seguela M, Inze D, Dissmeyer N, Li Y, Bevan MW (2017) Ubiquitylation activates a peptidase that promotes cleavage and destabilization of its activating E3 ligases and diverse growth regulatory proteins to limit cell proliferation in Arabidopsis.
Genes Dev. http://dx.doi.org/10.1101/gad.292235.116
Open Access
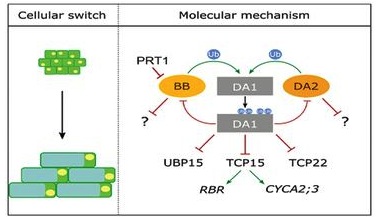
Mike Bevan (John Innes Centre) is the corresponding author of this study that also includes researchers from labs in Belgium, Germany and China. They investigate a fundamental determinant of organ shape, the pattern of cell proliferation that leads to final cell size. They show that two RING E3 ligases activate the DA1 peptidase that in turn affects the stabilization and activity of a range of other proteins including the transcription factors TEOSINTE BRANCED 1/CYCLOIDEA/PCF 15 (TCP15) and TCP22. Overall this results in continued cell proliferation and repression of endoreduplication, which ultimately serves to regulate the timing of the transition from cell proliferation to organ differentiation.
Mike discusses the science surrounding this paper on the GARNet YouTube channel.
Guo X,, Wang J, Gardner M, Fukuda H, Kondo Y, Etchells JP, Wang X, Mitchum MG. Identification of cyst nematode B-type CLE peptides and modulation of the vascular stem cell pathway for feeding cell formation. PLoS Pathog. http://dx.doi.org/10.1371/journal.ppat.1006142
Open Access
Peter Etchells (University of Durham) is a co-author on this US-led study that looks at the effect of nematode-delivered CLE-like peptides on cell growth and how that impacts parasitism. This study has identified a new class of peptides from nematodes that are similar to the plant B-type CLE-like peptide TDIF (tracheary element differentiation inhibitory factor). They show that the nematodes alter the activity of the TDIF-TDR (TDIF receptor)-WOX4 signaling module during infection, whose endogenous function acts during procambial meristem cell proliferation. A variety of mutants involved in this process show reduced infection and leading to the hypothesis that WOX4 is a potential target for nematode CLEs. When exogenous nematode CLE peptides are added to Arabidopsis roots they cause massive cell proliferation. This demonstrates that this response is clearly important for the establishment of nematode infection, usually in cambial cell files.
Bender KW, Blackburn RK, Monaghan J, Derbyshire P, Menke FL, Zipfel C, Goshe MB, Zielinski RE, Huber SC (2017) Autophosphorylation-based calcium (Ca2+) sensitivity priming and Ca2+/Calmodulin inhibition of Arabidopsis thaliana Ca2+-dependent protein kinase 28 (CPK28) J Biol Chem.
http://dx.doi.org/10.1074/jbc.M116.763243
Cyril Zipfel (The Sainsbury Lab) features for a second consecutive week on the Arabidopsis research roundup, this time as a co-author in a study that investigates the role of autophosphorylation in the regulation of calcium (Ca2+) dependent protein kinases (CPKs). In addition they evaluated the role of Calmodulin (CaM) on the activity of CPKs, something that had been previously overlooked. Indeed they show that CPK28 is a CaM-binding protein and that autophosphorylation causes increased activity, especially in low Ca2+ concentrations. Therefore this research provides a mechanistic insight into how a cell might respond to low levels of Ca2+.
Imprialou M, Kahles A, Steffen JG, Osborne EJ, Gan X, Lempe J, Bhomra A, Belfield E, Visscher A, Greenhalgh R, Harberd NP, Goram R, Hein J, Robert-Seilaniantz A, Jones J, Stegle O, Kover P, Tsiantis M, Nordborg M, Rätsch G, Clark RM, Mott R Genomic Rearrangements in Arabidopsis Considered as Quantitative Traits. Genetics. http://dx.doi.org/10.1534/genetics.116.192823
Open Access

Richard Mott (UCL) is corresponding author on this paper includes authors from throughout the UK, Europe and the US. They provide a new analysis of Arabidopsis populations that relies on the genome structural variation. They treat these structural variants as quantitative traits and subsequently map genetically in the same way as in a gene expression study. When a structural variant locus is linked to a genotype at a distant locus then it is designated as a site of transposition. Remarkably they show 25% of the structural variants can be assigned to the transposition events. This method of assessing structural variant loci is amendable to sequencing at low-coverage and this study identified loci that might be involved in germination and resistant to pathogens. Overall they find that genes within structural variants are more likely to be silenced and that this novel analysis technique is particularly useful when mapping transposition events.
Campbell L, Turner SR1(2017) A Comprehensive Analysis of RALF Proteins in Green Plants Suggests There Are Two Distinct Functional Groups. Front Plant Sci. http://dx.doi.org/10.3389/fpls.2017.00037
Open Access

This study from the lab of Simon Turner (University of Manchester) analyse Rapid Alkalinization Factor (RALFs) cysteine-rich peptides from across 51 plant species. They infer that these plant RALFs originate from four major clades in which the majority of the variation exists in the mature peptide sequence, indicative of clade-specific activities. Clade IV accounts for a third of the total peptides yet these lack a number of sequence features thought to be important for RALF function, which leads the authors to speculate that this clade should be thought of as containing RALF-related peptides instead of regular RALFs. Further experimental work is needed in order to define the true nature of the functional relationship between Clades I-III and Clade IV.
Ahmad L, Rylott EL, Bruce NC, Edwards R, Grogan G (2016) Structural evidence for Arabidopsis glutathione transferase AtGSTF2 functioning as a transporter of small organic ligands. FEBS Open Bio. http://dx.doi.org/10.1002/2211-5463.12168
Open Access
This paper links plant science and structural biology in a study that was undertaken at the University of York. Plant Glutathione transferases (GSTs) have multiple roles including in the detoxification of xenobiotics as well as in various non-catalytic roles. In this work the structure of the Arabidopsis AtGSTF2 is revealed in tandem with a variety of non-catalytic partners including indole-3-aldehyde, camalexin, the flavonoid quercetrin and its non-rhamnosylated analogue quercetin. These are thought to bind to AtGSTF2 by hydrophobic interactions at either one or two symmetrical binding sites. The authors speculate that this non-catalytic binding might have a possible role in ligand transport.

Podcast: Play in new window | Download